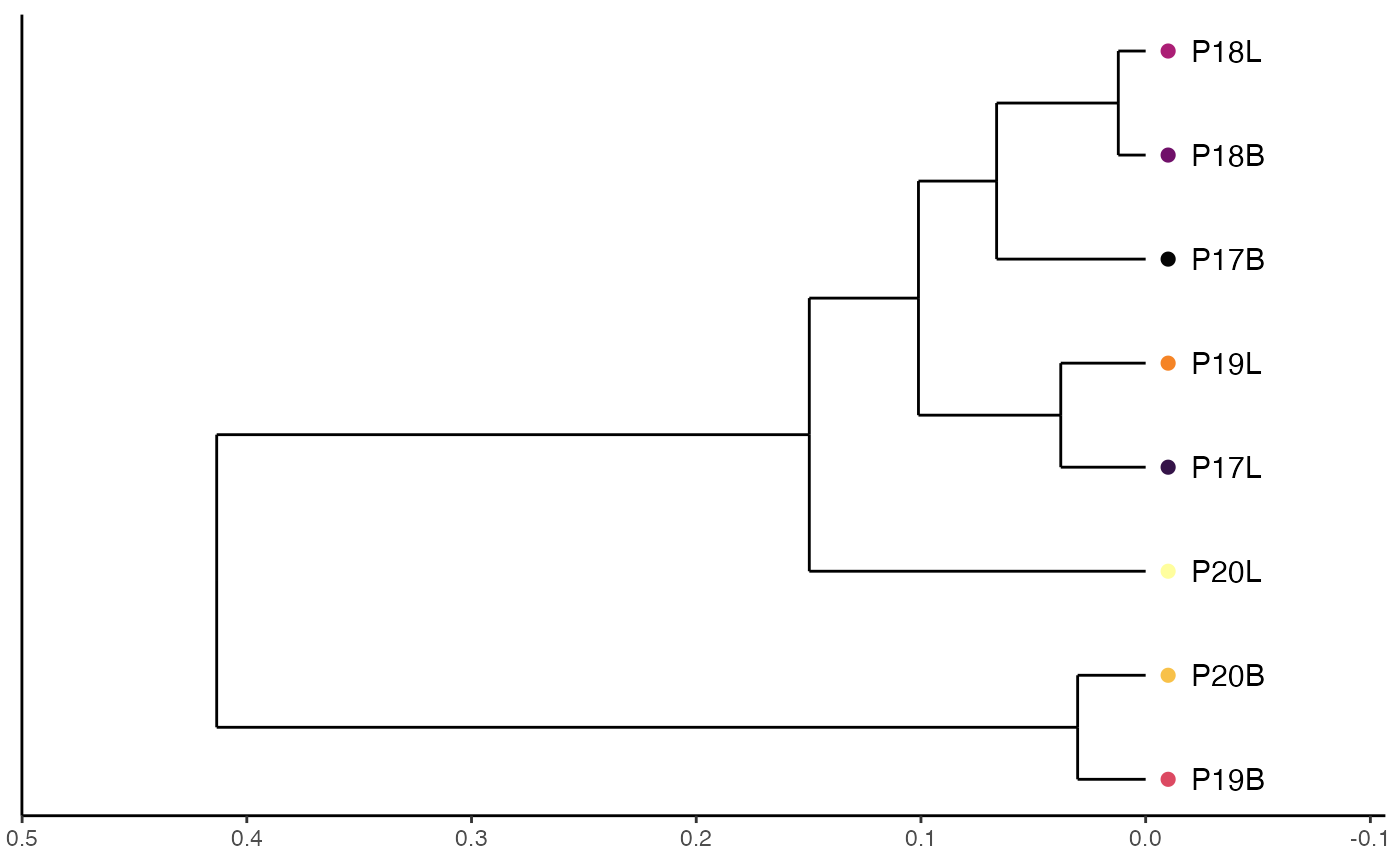
Plot powerTCR Clustering Based on Clonal Size
Source:R/clonalSizeDistribution.R
clonalSizeDistribution.RdThis function produces a hierarchical clustering of clones by sample using discrete gamma-GPD spliced threshold model. If using this model please read and cite powerTCR (more info available at PMID: 30485278).
Usage
clonalSizeDistribution(
input.data,
clone.call = NULL,
chain = "both",
method = "ward.D2",
threshold = 1,
group.by = NULL,
export.table = NULL,
palette = "inferno",
cloneCall = NULL,
exportTable = NULL,
...
)Arguments
- input.data
The product of
combineTCR(),combineBCR(), orcombineExpression().- clone.call
Defines the clonal sequence grouping. Accepted values are:
gene(VDJC genes),nt(CDR3 nucleotide sequence),aa(CDR3 amino acid sequence), orstrict(VDJC + nt). A custom column header can also be used.- chain
The TCR/BCR chain to use. Use
bothto include both chains (e.g., TRA/TRB). Accepted values:TRA,TRB,TRG,TRD,IGH,IGL,IGK,Light(for both light chains), orboth(for TRA/B and Heavy/Light).- method
The clustering parameter for the dendrogram.
- threshold
Numerical vector containing the thresholds the grid search was performed over.
- group.by
A column header in the metadata or lists to group the analysis by (e.g., "sample", "treatment"). If
NULL, data will be analyzed as by list element or active identity in the case of single-cell objects.- export.table
If
TRUE, returns a data frame or matrix of the results instead of a plot.- palette
Colors to use in visualization - input any hcl.pals.
- cloneCall
- exportTable
- ...
Additional arguments passed to the ggplot theme
Value
A ggplot object visualizing dendrogram of clonal size distribution
or a data.frame if exportTable = TRUE.
Details
The probability density function (pdf) for the Generalized Pareto Distribution (GPD) is given by: $$f(x|\mu, \sigma, \xi) = \frac{1}{\sigma} \left( 1 + \xi \left( \frac{x - \mu}{\sigma} \right) \right)^{-\left( \frac{1}{\xi} + 1 \right)}$$
Where:
\(\mu\) is a location parameter
\(\sigma > 0\) is a scale parameter
\(\xi\) is a shape parameter
\(x \ge \mu\) if \(\xi \ge 0\) and \(\mu \le x \le \mu - \sigma/\xi\) if \(\xi < 0\)
The probability density function (pdf) for the Gamma Distribution is given by: $$f(x|\alpha, \beta) = \frac{x^{\alpha-1} e^{-x/\beta}}{\beta^\alpha \Gamma(\alpha)}$$
Where:
\(\alpha > 0\) is the shape parameter
\(\beta > 0\) is the scale parameter
\(x \ge 0\)
\(\Gamma(\alpha)\) is the gamma function of \(\alpha\)
Examples
# Making combined contig data
combined <- combineTCR(contig_list,
samples = c("P17B", "P17L", "P18B", "P18L",
"P19B","P19L", "P20B", "P20L"))
# Using clonalSizeDistribution()
clonalSizeDistribution(combined,
clone.call = "strict",
method="ward.D2")