This functions allows for the calculation and visualizations of
various overlap metrics for clones. The methods include overlap
coefficient (overlap), Morisita's overlap index
(morisita), Jaccard index (jaccard), cosine
similarity (cosine) or the exact number of clonal
overlap (raw).
Usage
clonalOverlap(
input.data,
clone.call = NULL,
method = c("overlap", "morisita", "jaccard", "cosine", "raw"),
chain = "both",
group.by = NULL,
order.by = NULL,
export.table = NULL,
palette = "inferno",
cloneCall = NULL,
exportTable = NULL,
...
)Arguments
- input.data
The product of
combineTCR(),combineBCR(), orcombineExpression()- clone.call
Defines the clonal sequence grouping. Accepted values are:
gene(VDJC genes),nt(CDR3 nucleotide sequence),aa(CDR3 amino acid sequence), orstrict(VDJC + nt). A custom column header can also be used.- method
The method to calculate the
overlap,morisita,jaccard,cosineindices orrawfor the base numbers- chain
The TCR/BCR chain to use. Use
bothto include both chains (e.g., TRA/TRB). Accepted values:TRA,TRB,TRG,TRD,IGH,IGL,IGK,Light(for both light chains), orboth(for TRA/B and Heavy/Light).- group.by
A column header in the metadata or lists to group the analysis by (e.g., "sample", "treatment"). If
NULL, data will be analyzed by list element or active identity in the case of single-cell objects.- order.by
A character vector defining the desired order of elements of the
group.byvariable. Alternatively, usealphanumericto sort groups automatically.- export.table
If
TRUE, returns a data frame or matrix of the results instead of a plot.- palette
Colors to use in visualization - input any hcl.pals
- cloneCall
- exportTable
- ...
Additional arguments passed to the ggplot theme
Details
The formulas for the indices are as follows:
Overlap Coefficient: $$overlap = \frac{\sum \min(a, b)}{\min(\sum a, \sum b)}$$
Raw Count Overlap: $$raw = \sum \min(a, b)$$
Morisita Index: $$morisita = \frac{\sum a b}{(\sum a)(\sum b)}$$
Jaccard Index: $$jaccard = \frac{\sum \min(a, b)}{\sum a + \sum b - \sum \min(a, b)}$$
Cosine Similarity: $$cosine = \frac{\sum a b}{\sqrt{(\sum a^2)(\sum b^2)}}$$
Where:
\(a\) and \(b\) are the abundances of species \(i\) in groups A and B, respectively.
Examples
# Making combined contig data
combined <- combineTCR(contig_list,
samples = c("P17B", "P17L", "P18B", "P18L",
"P19B","P19L", "P20B", "P20L"))
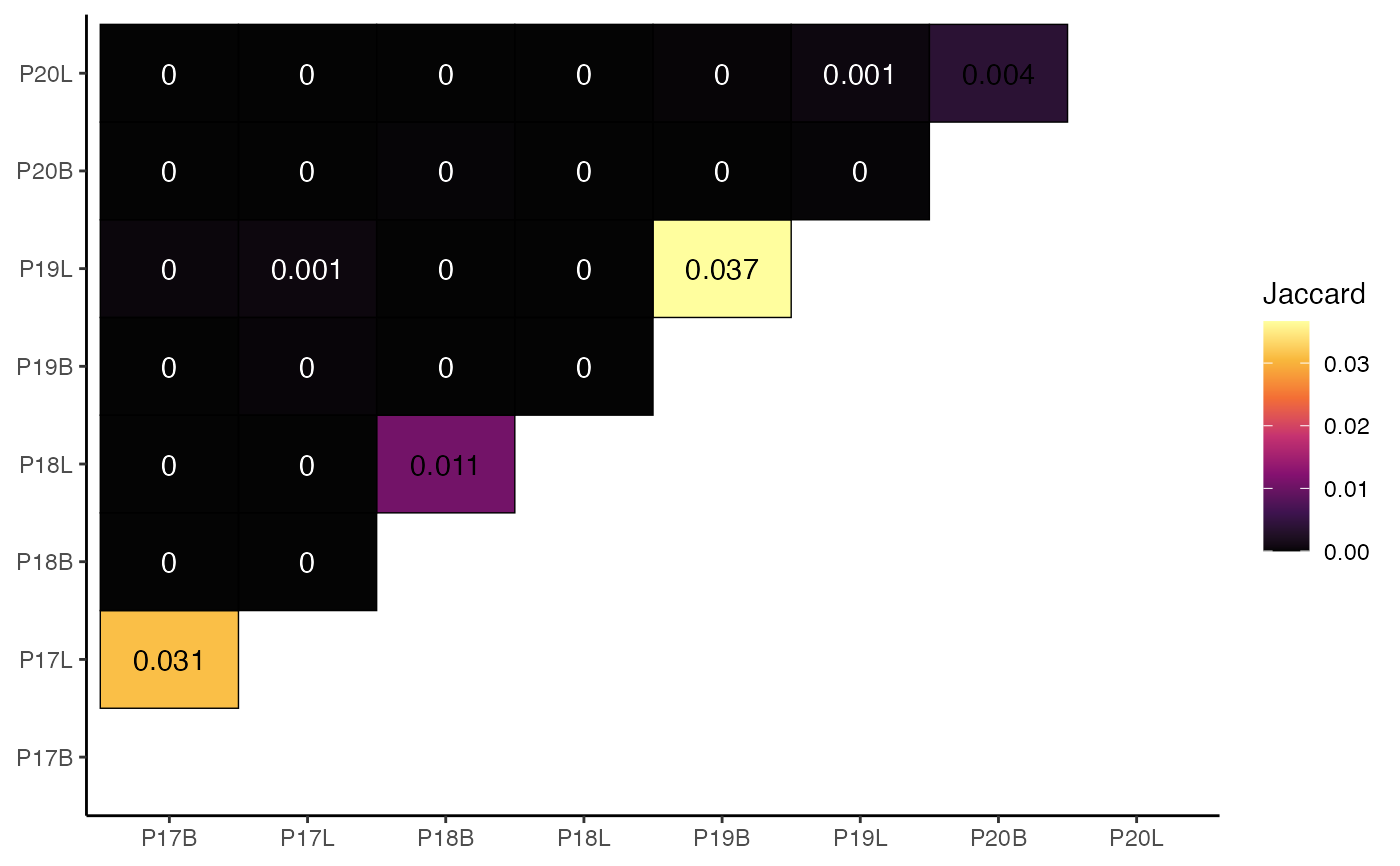
# Using clonalOverlap()
clonalOverlap(combined,
clone.call = "aa",
method = "jaccard")